Today, there are only a few therapies approved to treat COVID-19, but while novel therapies can take decades and billions of dollars to develop, are there opportunities to repurpose existing drugs for new therapies? Our latest whitepaper showcases how CAS Knowledge Graphs reveal new connections and insights that identify drugs to potentially repurpose.
Drug repurposing is critical for faster development of therapies. However, assembling all the critical information and connections around new proteins, viruses, targets, pathways, and clinical information can be challenging. This demonstrates how CAS Knowledge Graphs can identify top clinical candidates to repurpose for COVID-19 therapies.
A knowledge graph combines data from disparate sources to model a particular area. It describes data in nodes and edges. Nodes represent each point of data and edges represent the relationship between them. The image below provides a simplified example of a knowledge graph that predicts which drugs might inhibit vascular inflammation.
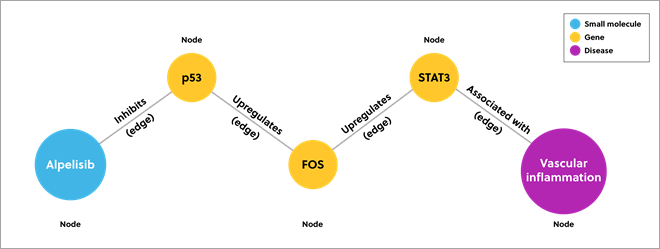
Figure 1. Example of a knowledge graph showing the connections between data using nodes and edges
Traditional databases may only show direct connections (direct inhibitors of transcription factor STAT3), but a knowledge graph can show deeper data connections. In this example, the knowledge graph presents inhibitors that act further down the pathway.
The CAS Biomedical Knowledge Graph combines human-curated data from the CAS Content CollectionTM with publicly available biomedical data.
It contains high-quality data from over 6 million small molecules, 24,000 diseases, and 26,000 human and viral genes. A knowledge graph reveals insights that would not be possible using traditional research methods.
Our approach included two core components to uncover potential drug candidates for COVID-19:
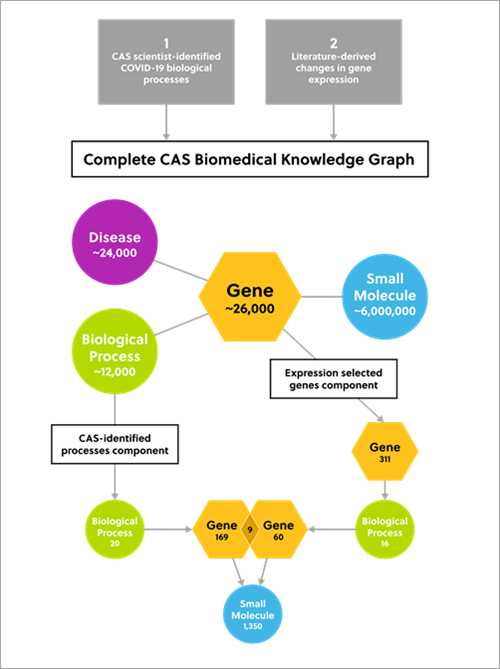
Figure 2. Diagram outlining the two-component approach to identify potential small molecule drug candidates for COVID-19 therapeutics
Using the knowledge graph, we identified:
The analysis identified 1,350 small molecules that could offer potential for repurposing as therapeutics for COVID-19.
Once we identified potential molecules, we assessed the power of their connections and boosted scores accordingly. To do this, we used a novel algorithmic method to rank each molecule. The equation evaluated the relationships between the small molecules and the interactions with the genes and biological processes identified in our two-component approach.
For example, a cytokine storm was considered an important connection. We then evaluated the relationships between the small molecules and the interactions with the genes and biological processes identified in our two-component approach. Score boosts were given to important connections, such as to cytokine storm and to small molecules that have an activating relationship with genes, given the rarity of these occurrences.
Thus, we were able to develop a ranking table of all the small molecules and we present the top 50 in our whitepaper. In Figure 2 below, you can see the top 10 scoring drug candidates from the results. The size of the node corresponds to the number of connections to other nodes.
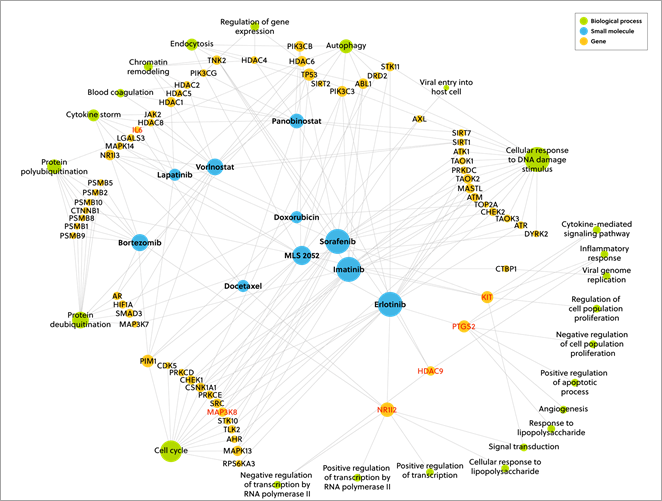
Figure 3. A network diagram showing the connection of the top 10 scoring drug candidates from the results with the size of the nodes corresponds to the number of connections to other nodes
Out of the top 50 drugs identified in our ranking table, 11 are currently in clinical trials for treating COVID-19. This provides validation of our results.
Our biomedical knowledge graph uncovers four drug classes that have been linked previously to SARS-CoV-2 or general viral infection mechanisms. The four drug classes include:
Kinase inhibitors
These were the single largest class of drugs found in our results. Kinases are involved in almost all biological processes and their activities are dysregulated in many diseases. Receptor tyrosine kinases (RTKs) are involved in the cell entry of many viruses. The kinase inhibitors identified included those affecting RTKs such as EGF, FGF, PDGF, and ALK receptors, as well as non-receptor tyrosine kinases such as Bruton tyrosine kinase. Serine-threonine kinase inhibitors targeting receptors B-RAF, PKC, PIM, and GSK-2beta were also identified by our knowledge graph.
Histone deacetylase inhibitors (HDIs )
HDIs regulate gene expression by reducing histone deacetylation. HDIs reduce the expression of both angiotensin-converting enzyme 2 (ACE2), the main cell surface receptor of SARS-CoV-2, and the ABO glycosyltransferase, an enzyme that helps regulate blood type, which is a known COVID-19 risk factor. HDIs also regulates several of the chemokines and cytokines involved in the immune response in COVID-19. As such, their inclusion in the results is logical.
Microtubule-regulating agents
Microtubules are filaments composed of tubulin subunits. Studies have shown that SARS-CoV-2 proteins interact with microtubules or microtubules-associated proteins. Our results uncovered that microtubule-regulating agents, such as docetaxel, colchicine, and mebendazole, may be of use in disrupting SARS-CoV-2 infection. Colchicine is already in clinical trials for the treatment of COVID-19 patients.
Protease inhibitors
Of the protease inhibitors identified, most were proteasome inhibitors. Studies have shown that the ubiquitin-proteasome system is involved in viral replication and the cytokine storm, including in diseases associated with coronavirus. Protease inhibitors are a logical choice for exploring in relation to COVID-19. Indeed, several such inhibitors are already being investigated as COVID-19 therapeutics. Some were found in our results, such as bortezomib, carfilzomib, and saxagliptin.
The methodology behind our knowledge graph enhances potential drug identification for COVID-19 treatment and will be of great value for drug discovery in other diseases beyond COVID-19, such as Alzheimer’s disease, Parkinson’s disease, autoimmune diseases, cancer, and even rare diseases. Our knowledge graphs are both scalable and modular and offer great value to all areas of science, including chemistry, nutrition, and renewable energies. The opportunities are vast.
Read our whitepaper to find out more about our methodologies and results, and how we harness the power of the CAS Biomedical Knowledge Graph to identify potential therapeutics for COVID-19.